Computer vision deep learning examples based on medical imaging
Let's explore a detection problem in medical imaging based on a public dataset. We'll use MLFlow's "Projects" functionality, in which a self-contained modeling module is trained, has its performance logged in MLFlow, and is able to be checked out as a deployable model image. There's a configurable implementation of this in my aganse/py_tf2_gpu_dock_mlflow repo.
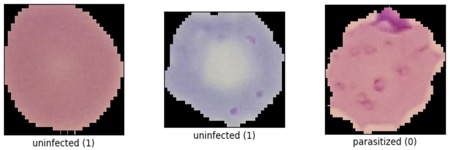
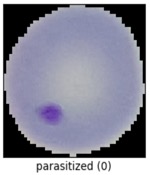
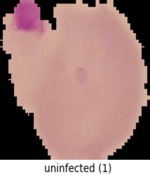
We'll use the malaria detection dataset from the Tensorflow datasets, which contains a balanced, labeled dataset of about 27,000 thin blood smear slide images of cells, and let's see how well we can detect malaria parasite presence in the images. This dataset is used to train/test different variations of image classification models, including VGG-16 and various sizes of more basic convolutional networks.
The Python/Tensorflow2 setup in aganse/py_tf2_gpu_dock_mlflow uses MLflow with GPU runs in a Docker container to train, evaluate, and log models to MLflow's model registry. Logged models can then be easily served via REST API or downloaded from the registry into python code or their own new Docker image for further use. The training computation itself is handled entirely in the container; for that the host system only needs the Nvidia driver and Docker installed. (However, currently to kick off the training you do still need a Python environment with MLflow installed.). The training script includes function definitions for the neural network specification, so models are easily experimented with independently of the rest of the code.
The MLflow screenshot here gives an idea of the information that gets logged with the runs. If you don't have an MLflow instance running already, you can pop one up in minutes with my aganse/docker_mlflow_db implementation. And aganse/py_tf2_gpu_dock_mlflow provides instructions for quickly cranking up both modeling and MLlflow resources in an AWS EC2 instance if desired.
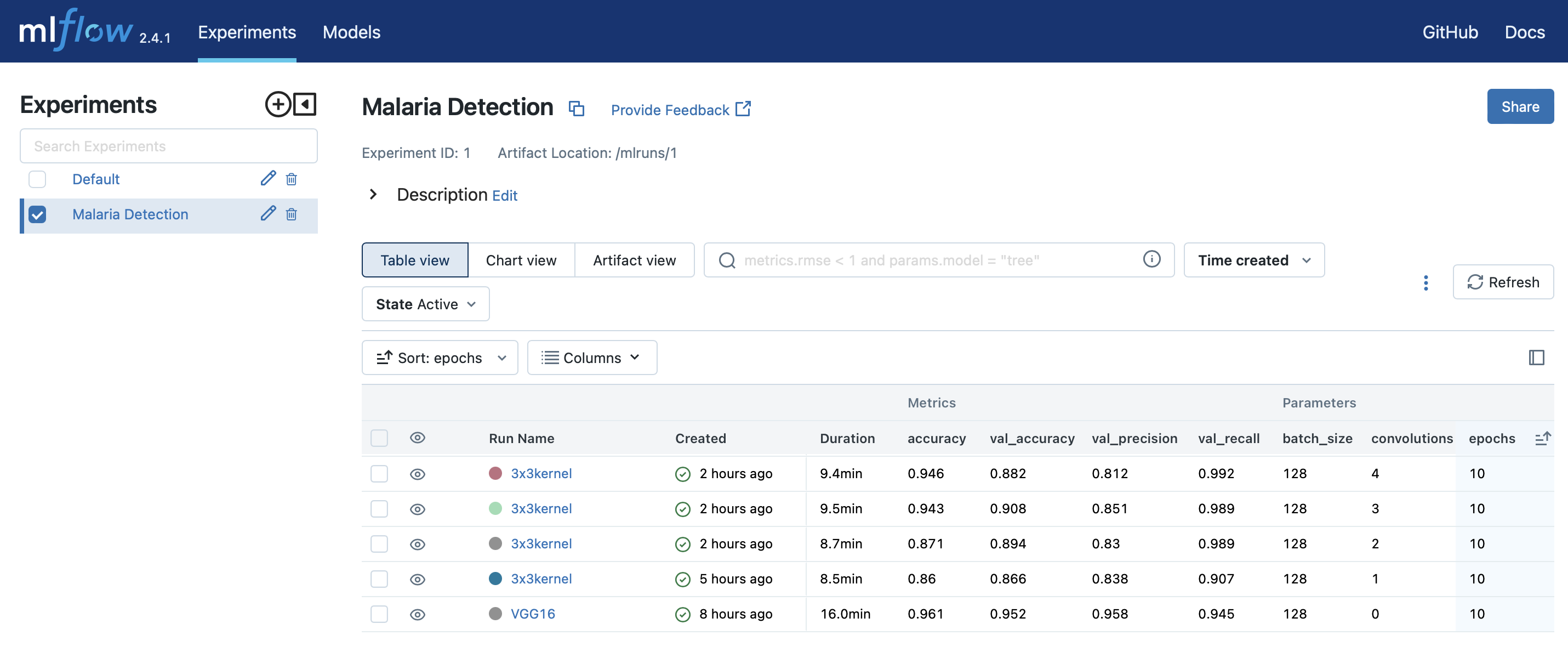
You can see that of the 5 runs in the list, the VGG16 run certainly performs the best; but of course that's traded off with being a much bigger model (like over 1000x more parameters!). So on one hand, one could very likely find some much smaller "hand-made" model that works as well as VGG16 does here. But on other hand, VGG16 is all ready to go in Keras and it's as easy to implement as just



Let's explore a detection problem in medical imaging based on a public dataset. We'll use MLFlow's "Projects" functionality, in which a self-contained modeling module is trained, has its performance logged in MLFlow, and is able to be checked out as a deployable model image. There's a configurable implementation of this in my aganse/py_tf2_gpu_dock_mlflow repo.
We'll use the malaria detection dataset from the Tensorflow datasets, which contains a balanced, labeled dataset of about 27,000 thin blood smear slide images of cells, and let's see how well we can detect malaria parasite presence in the images. This dataset is used to train/test different variations of image classification models, including VGG-16 and various sizes of more basic convolutional networks.
The Python/Tensorflow2 setup in aganse/py_tf2_gpu_dock_mlflow uses MLflow with GPU runs in a Docker container to train, evaluate, and log models to MLflow's model registry. Logged models can then be easily served via REST API or downloaded from the registry into python code or their own new Docker image for further use. The training computation itself is handled entirely in the container; for that the host system only needs the Nvidia driver and Docker installed. (However, currently to kick off the training you do still need a Python environment with MLflow installed.). The training script includes function definitions for the neural network specification, so models are easily experimented with independently of the rest of the code.
The MLflow screenshot here gives an idea of the information that gets logged with the runs. If you don't have an MLflow instance running already, you can pop one up in minutes with my aganse/docker_mlflow_db implementation. And aganse/py_tf2_gpu_dock_mlflow provides instructions for quickly cranking up both modeling and MLlflow resources in an AWS EC2 instance if desired.

You can see that of the 5 runs in the list, the VGG16 run certainly performs the best; but of course that's traded off with being a much bigger model (like over 1000x more parameters!). So on one hand, one could very likely find some much smaller "hand-made" model that works as well as VGG16 does here. But on other hand, VGG16 is all ready to go in Keras and it's as easy to implement as just
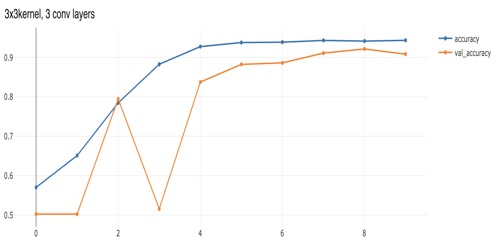
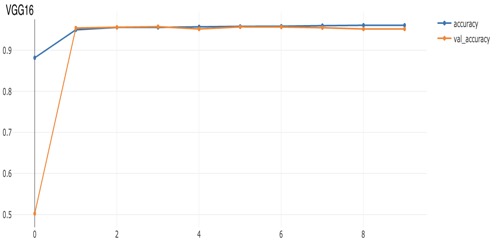
model.add(VGG16(input_shape=IMAGE_SHAPE, weights="imagenet")). We can see in comparing the performance plots below that VGG16 converged much more quickly than model with just a few convolutional layers, but also ended up with slightly higher accuracy. In those plots the "val_accuracy" is the validation-data accuracy while "accuracy" is the training-data accuracy. The dataset here is balanced, with plenty images of both positives and negatives, so accuracy works for our purposes here, but often in problems where we're detecting something rare (often resulting in heavily unbalanced training data), we'd need to focus on other metrics such as F1 or balanced accuracy.

